Tutorial 3: Synthetic Aorta Pipeline
Goal: Run the complete AortaCFD pipeline on a patient-like synthetic geometry -- an aortic arch with coarctation and three supra-aortic branches.
 |
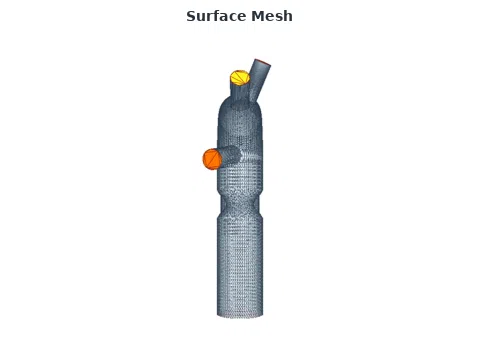 |
|---|---|
| Geometry: arch + branches + coarctation | Mesh: surface → cross-section (200K cells) |
Hands-On Exercise
Download: exercise_g3_quick.tar.gz (523 KB) | ~30 sec | OpenFOAM 12
Synthetic aortic arch (G3) with 65% coarctation. Complete pipeline from STL to solution.
Introduction
The previous tutorials introduced meshing and boundary conditions on simple tube geometries. This tutorial brings those concepts together on a patient-like aortic arch -- a synthetic geometry generated by the Blender parametric tool in this project. The G3 geometry includes an aortic arch with three supra-aortic branches and a 65% coarctation (area reduction at the aortic isthmus), making it a clinically relevant benchmark.
Synthetic geometries serve an essential role in computational haemodynamics research:
- Controlled parameters: The coarctation severity, branch angles, and vessel diameters are known exactly, enabling systematic parametric studies.
- Reproducibility: Any researcher can regenerate the identical geometry from the Blender script, eliminating segmentation variability.
- Ground truth: Because the geometry is analytically defined, mesh convergence studies and code verification are straightforward.
- No ethics approval required: Synthetic cases can be shared freely as teaching and benchmarking material.
By the end of this tutorial you will be able to:
- Identify the STL patches and their anatomical correspondence.
- Read and interpret an AortaCFD configuration file.
- Run the full pipeline (mesh generation through solver execution).
- Examine the output directory structure and verify mesh quality.
- Understand the haemodynamic significance of aortic coarctation.
The G3 Geometry
The G3 synthetic aortic arch has the following specifications:
| Feature | Value |
|---|---|
| Main aortic diameter | 24 mm |
| Coarctation severity | 65% area reduction |
| Coarctation location | Aortic isthmus (distal to left subclavian) |
| Supra-aortic branches | 3 (brachiocephalic, LCCA, LSA) |
| Outlets | 4 (3 branches + descending aorta) |
| STL units | Millimetres |
The coarctation creates a geometric narrowing at the isthmus -- the segment of the aorta just distal to the left subclavian artery origin. Clinically, coarctation of the aorta is a congenital condition that produces a trans-stenotic pressure gradient and accelerated flow through the narrowing. The ESC guidelines define a resting pressure gradient exceeding 20 mmHg as the threshold for intervention Erbel 2014.
Anatomical Orientation
The aortic arch curves from the ascending aorta (rising from the left ventricle) over the pulmonary trunk and descends as the thoracic aorta. Three branches arise from the superior aspect of the arch:
- Brachiocephalic trunk (innominate artery) -- supplies the right arm and right side of the head.
- Left common carotid artery (LCCA) -- supplies the left side of the head.
- Left subclavian artery (LSA) -- supplies the left arm.
Case Input Structure
Extract the exercise and examine the contents:
The input directory contains the following files:
| File | Purpose | Anatomical Correspondence |
|---|---|---|
inlet.stl |
Inlet patch surface | Ascending aorta (flow enters here) |
outlet1.stl |
Outlet patch 1 | Brachiocephalic trunk |
outlet2.stl |
Outlet patch 2 | Left common carotid artery |
outlet3.stl |
Outlet patch 3 | Left subclavian artery |
outlet4.stl |
Outlet patch 4 | Descending aorta |
wall_aorta.stl |
Wall patch surface | Aortic wall (WSS computed here) |
config.json |
Simulation configuration | All pipeline parameters |
Each STL file represents one boundary patch of the simulation domain. The
patches must be watertight (no gaps between adjacent patches) and oriented
with outward-facing normals. AortaCFD reads the inlet diameter from
inlet.stl to determine the mesh resolution.
STL Naming Convention
AortaCFD identifies patches by filename: inlet.stl is always the
inlet, files matching outlet*.stl are outlets, and wall_aorta.stl
is the vessel wall. This convention must be followed for the pipeline
to assign boundary conditions correctly.
Understanding the Config
Open the configuration file:
The configuration is organised into sections. The quick tutorial config uses deliberately simplified settings for fast execution:
case_info
Identifies the case. The case_name is used to create the output directory.
physics
| Parameter | Value | Notes |
|---|---|---|
model |
laminar | No turbulence model; appropriate for Re < 4000 |
density |
1060 kg/m^3 | Standard blood density |
dynamic_viscosity |
0.004 Pa.s | Newtonian blood assumption |
Kinematic Viscosity
OpenFOAM internally uses kinematic viscosity: \(\nu = \mu / \rho = 0.004 / 1060 = 3.774 \times 10^{-6}\) m^2/s. AortaCFD performs this conversion automatically.
numerics
The quick config uses the robust profile (first-order schemes) for
maximum stability on the coarse mesh. For production simulations, switch to
standard.
mesh
| Parameter | Value | Notes |
|---|---|---|
cells_per_diameter |
10 | Coarse; produces ~50,000--100,000 cells |
boundary_layers |
disabled | No prismatic layers; WSS will be approximate |
Boundary Layers Disabled
This quick config intentionally disables boundary layers to reduce meshing time. Wall shear stress computed without boundary layers is unreliable. For any study reporting WSS metrics, enable boundary layers with at least 3--5 prismatic layers.
geometry
The scale_factor of 0.001 converts STL coordinates from millimetres to
metres (OpenFOAM's internal unit system). This is a critical parameter --
an incorrect scale factor is one of the most common sources of error in
cardiovascular CFD.
Always Verify the Scale Factor
If the STL is already in metres, set scale_factor to 1.0. If it is in
millimetres (as is standard for medical imaging STL exports), use 0.001.
A wrong scale factor produces a domain 1000x too large or too small,
leading to nonsensical velocities and pressures. Check the inlet audit
report after case setup to verify the inlet area is physiologically
reasonable (~3--5 cm^2 for an adult aorta).
boundary_conditions
{
"boundary_conditions": {
"inlet": {
"type": "CONSTANT",
"cardiac_output": 5.0,
"profile": "plug"
},
"outlets": {
"type": "zeroGradient"
}
}
}
The quick config uses the simplest boundary conditions: constant inlet flow
and zeroGradient outlets. This is sufficient for verifying that the mesh and
solver work correctly. For physiological results, use TIMEVARYING inlet
and 3EWINDKESSEL outlets as described in
Tutorial 2.
simulation_control
| Parameter | Value | Notes |
|---|---|---|
end_time |
0.05 s | Very short; just enough to see the flow develop |
parallel |
false | Serial execution; no domain decomposition |
num_processors |
1 | Single core |
Production Settings
A full cardiac cycle simulation (3 cycles at 0.8 s each = 2.4 s) with Windkessel outlets, cpd = 15, boundary layers, and 8 processors is a typical production configuration. The quick config here runs in ~30 seconds to demonstrate the pipeline workflow.
Running the Pipeline
There are two ways to execute the simulation.
Option A: Using AortaCFD (Recommended)
If you have AortaCFD cloned and its virtual environment set up:
source /opt/openfoam12/etc/bashrc
cd /path/to/AortaCFD
source venv/bin/activate
python run_patient.py G3 \
--config /path/to/exercise_g3_quick/config.json \
--run-name tutorial \
--steps case,mesh,boundary,solver
AortaCFD orchestrates the complete pipeline:
- case -- Creates the OpenFOAM directory structure, copies STL files, renders Jinja2 templates for all dictionaries.
- mesh -- Runs
blockMesh,surfaceFeatures,snappyHexMesh, andcheckMesh. - boundary -- Generates the
0/Uand0/pboundary condition files from the configuration. - solver -- Executes
foamRun(ormpirun -np N foamRunfor parallel cases).
Option B: Manual OpenFOAM Steps
If you prefer to run the OpenFOAM utilities directly (useful for learning what AortaCFD automates):
source /opt/openfoam12/etc/bashrc
cd exercise_g3_quick
# 1. Generate background mesh
blockMesh
# 2. Extract surface features
surfaceFeatures
# 3. Generate volume mesh (castellate, snap, layers)
snappyHexMesh -overwrite
# 4. Check mesh quality
checkMesh
# 5. Run the solver
foamRun
The -overwrite Flag
By default, snappyHexMesh writes each stage to a separate time directory
(1/, 2/, 3/). The -overwrite flag writes the final mesh directly to
constant/polyMesh/, which is the expected location for subsequent
utilities and the solver.
Using the Provided run.sh Script
The exercise includes a convenience script:
This script sources the OpenFOAM environment, runs all meshing steps, checks the mesh, and starts the solver. It is equivalent to Option B above.
Examining the Output
Mesh Quality
After meshing, inspect the checkMesh output:
Expected output for cpd = 10 on the G3 geometry:
Checking geometry...
Overall domain bounding box (-0.015 -0.06 -0.015) (0.015 0.005 0.04)
Mesh has 3 geometric (non-empty/wedge) directions (1 1 1)
Mesh has 3 solution (non-empty) directions (1 1 1)
Checking topology...
Cell to face addressing OK.
Point usage OK.
Checking geometry...
Max non-orthogonality = 58.2 OK.
Max skewness = 2.1 OK.
Max aspect ratio = 18.7 OK.
Mesh OK.
Cell Count
With cpd = 10 and no boundary layers, expect approximately 50,000 to
100,000 cells. The exact count depends on the snappyHexMesh refinement
algorithm's interaction with the geometry. Check the checkMesh output
for the line reporting cells:.
Solution Fields
After the solver completes, time directories appear in the case:
Each time directory contains the velocity (U) and pressure (p) fields.
Open the final time step in ParaView:
- Click Apply to load the mesh.
- Set representation to Surface.
- Colour by
U(velocity magnitude) -- observe the flow acceleration through the coarctation. - Colour by
p(pressure) -- observe the pressure drop across the narrowing.
Key Flow Features
On the G3 geometry with 65% coarctation, you should observe:
| Feature | Location | Physical Significance |
|---|---|---|
| Accelerated jet | Through the coarctation | Continuity: reduced area forces higher velocity |
| Pressure drop | Across the coarctation | Energy loss due to acceleration and viscous dissipation |
| Recirculation zone | Distal to the coarctation | Flow separation from the expansion |
| Branch flow | Supra-aortic arteries | Flow diverted to head and arms |
The Coarctation Jet
The 65% area reduction forces the blood to accelerate through the narrowing. By the continuity equation, if the area is reduced by 65%, the velocity increases by a factor of approximately 2.9. This high-velocity jet impinges on the distal wall, creating elevated wall shear stress, and the sudden expansion downstream produces a recirculation zone with disturbed flow. These are the primary haemodynamic features of aortic coarctation that guide clinical intervention decisions.
Exercise: Modify the Config and Re-Run
-
Copy the configuration file:
-
Edit
config_standard.jsonto change:numerics.profilefrom"robust"to"standard"mesh.cells_per_diameterfrom10to15mesh.boundary_layers.enabledfromfalsetotrue
-
Re-run the pipeline with the updated config.
-
Compare the two results:
- How does the cell count change?
- Is the
checkMeshquality different? - Are the velocity gradients near the coarctation sharper with the standard profile?
Expected Differences
Increasing cpd from 10 to 15 approximately doubles the cell count (cubic scaling). Enabling boundary layers adds prismatic cells near the wall, improving WSS resolution. The standard profile resolves sharper velocity gradients through the coarctation compared to the robust profile.
Summary
| Concept | Key Takeaway |
|---|---|
| Synthetic geometry | Controlled, reproducible, known ground truth; ideal for benchmarking |
| G3 coarctation | 65% area reduction at the isthmus; creates pressure gradient and jet |
| STL patches | inlet, outlet1--4, wall_aorta; named by convention for AortaCFD |
| scale_factor | 0.001 converts mm to m; verify via inlet area in audit report |
| Quick config | cpd = 10, robust, no layers, constant inlet, 0.05 s; ~30 seconds |
| Pipeline steps | case, mesh, boundary, solver; can run individually or together |
| Output | Time directories with U and p fields; checkMesh for quality |
What's Next
In Tutorial 4: Hemodynamics and Post-Processing, you will use pre-computed pulsatile results on this same G3 geometry to compute and interpret clinically relevant haemodynamic metrics: TAWSS, OSI, RRT, and trans-coarctation pressure gradient.
References
Full bibliography on the References page.
Found an issue or have a suggestion for this page?
Open a GitHub issue